EEG Software That Actually Works for Researchers
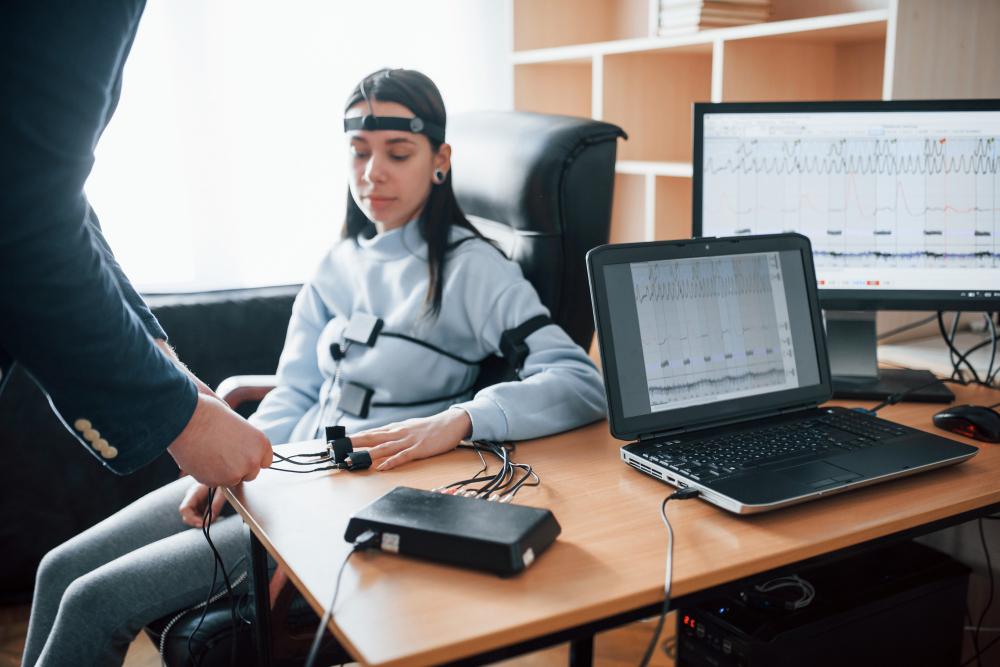

The Data Is Only as Good as the Tools You Use to Read It
You've spent hours setting up your montage, prepping your subjects, running your protocol. The EEG data is collected. And now comes the part that either validates all that effort or quietly undermines it — processing, analyzing, and interpreting what you actually captured.
This is where EEG software becomes the real center of the operation. Not the headset. Not the amplifier. The software.
And yet, it's the piece that researchers — especially those newer to the field — spend the least time thinking about before committing to a workflow. They inherit whatever their lab uses, or they pick the most popular name they've heard in a seminar, and they run with it. That's understandable. But it's also a decision that shapes everything downstream: your artifact rejection methods, your frequency analysis approach, your source localization capability, your ability to collaborate with other labs, and ultimately the reproducibility of your findings.
Let's slow down and actually think through this.
What EEG Software Is Really Being Asked to Do
Before comparing tools, it helps to be clear on the scope of what a capable EEG software environment needs to handle. Most researchers know the basics — filtering, epoching, artifact rejection. But modern EEG analysis is substantially more complex than that.
Signal preprocessing
Raw EEG is messy. Muscle artifacts, eye blinks, electrical interference, motion noise — the preprocessing pipeline is where you clean the signal before you can trust your analysis. The sophistication of a platform's preprocessing tools, and the degree of control it gives you over each step, matters enormously. Automated pipelines are convenient, but they can also quietly introduce decisions you haven't reviewed.
Time-frequency analysis
Whether you're working with event-related potentials, oscillatory dynamics, or connectivity patterns, your software needs to handle time-frequency decomposition reliably — Fourier-based, wavelet-based, Hilbert transform approaches. Understanding what your tool is doing under the hood, not just what button to click, is part of doing defensible science.
Independent component analysis
ICA has become a standard method for separating brain signal from artifacts like eye movements and muscle noise. Good EEG software gives you real control over ICA — component visualization, classification tools, and the ability to make informed decisions about what to remove rather than delegating that entirely to automation.
Connectivity and source analysis
For research pushing into network-level questions — how different brain regions communicate, where activity originates — source localization and connectivity analysis become essential. Not every platform handles these equally well, and the assumptions embedded in different approaches can lead to meaningfully different results.
Spike detection and event marking
In clinical contexts especially, automated detection of events in the EEG — epileptiform discharges, sleep spindles, sharp waves — is critical. EEG spike detection capabilities vary widely across platforms, from basic threshold algorithms to sophisticated machine learning classifiers. If this is part of your workflow, it deserves specific evaluation, not an afterthought.
The Landscape of EEG Software Options in the US
The US research and clinical community has access to a range of EEG software platforms, each with distinct strengths, communities, and learning curves.
EEGLAB
One of the most widely used open-source platforms in academic neuroscience. Built on MATLAB, EEGLAB has an enormous plugin ecosystem and a decades-long track record in research settings. Its flexibility is a major advantage — if you can code, you can extend it almost indefinitely. The learning curve is real, but the community support is extensive.
MNE-Python
For labs moving toward Python-based workflows, MNE-Python has become the dominant choice. It's genuinely powerful, actively maintained, and increasingly well-documented. The shift away from MATLAB dependency has made it more accessible for many researchers, especially trainees who learned Python before ever touched MATLAB.
Brainstorm
A MATLAB-based platform with a graphical interface that feels more approachable than EEGLAB for many users. Brainstorm has strong MEG and EEG support, solid source localization tools, and is particularly popular in clinical research and educational contexts.
BESA and other commercial platforms
Commercial platforms like BESA offer deep functionality, strong technical support, and are common in clinical settings where reliability and compliance matter. The tradeoff is cost and the closed nature of the codebase.
Cloud-based and collaborative platforms
A newer category that's growing fast. Platforms designed for collaborative analysis, reproducible pipelines, and large-scale data sharing are increasingly important as neuroscience moves toward open science practices. Neuromatch is a notable name in this space — contributing to education, open-source tooling, and community-building around computational neuroscience in ways that are reshaping how the field trains and connects researchers.
The Open Science Question
This is increasingly unavoidable in US neuroscience. Funders — including NIH — are pushing hard for data sharing, pre-registration, and reproducible workflows. The EEG software you choose has direct implications for all three.
Open-source platforms like MNE-Python and EEGLAB allow full transparency in your processing pipeline. Every step can be scripted, documented, version-controlled, and shared alongside your data and findings. Reviewers can see exactly what you did. Other labs can reproduce it.
Commercial platforms are improving on this front, but by nature, a proprietary codebase limits transparency in ways that open-source doesn't. If reproducibility and open science compliance are priorities — and increasingly they should be — this factor deserves weight in your decision.
Choosing EEG Software: A Framework for Making the Decision Well
Rather than recommending a single platform as universally superior, here's a framework for thinking through the decision in a way that actually fits your context.
Start with your research questions
What are you actually trying to understand? ERP components, oscillatory dynamics, connectivity, source localization, clinical event detection? The analysis methods your questions require should drive software selection — not the reverse.
Assess your team's technical capacity
A platform that requires extensive custom coding to use effectively is a liability if your lab doesn't have coding expertise. Conversely, an overly simplified GUI may constrain sophisticated analysis for experienced researchers. Be honest about where your team actually is.
Consider your collaboration network
If your primary collaborators all run MNE-Python pipelines, being the EEGLAB holdout creates friction. Interoperability and shared workflows reduce friction and improve collaboration. The community your platform belongs to matters.
Think about your data volume and computing needs
High-density EEG, long recordings, large cohorts — these create computational demands that not all platforms handle gracefully. Cloud-based solutions and well-optimized Python tools tend to scale better than legacy MATLAB-based environments for large datasets.
Factor in training and onboarding
How will new lab members learn the workflow? Is there documentation? Are there courses? Is there a community that answers questions? Sustainability of your pipeline depends on the ability to onboard people without starting from scratch every time.
The Clinical Context: Different Priorities, Different Needs
It's worth separating the research and clinical contexts because the priorities genuinely differ.
In clinical EEG — epilepsy monitoring, ICU surveillance, sleep studies — reliability, regulatory compliance, and interpretability are paramount. Automated detection features need to be validated and auditable. The workflow needs to work for clinicians who aren't computational neuroscientists.
In this context, eeg software selection often tilts toward platforms with strong clinical track records, robust technical support, and features purpose-built for clinical interpretation — even if that means sacrificing some of the flexibility that research environments value.
The good news is that the gap between clinical and research tools is narrowing. Platforms are increasingly designed to serve both communities, and the cross-pollination of methods — clinical-grade spike detection informing research, research-grade source analysis informing clinical interpretation — is genuinely enriching both fields.
Don't Let Your Software Choice Be an Afterthought
The EEG signal you've worked so hard to collect deserves processing and analysis tools that match the rigor of your data collection. Whether you're running a cognitive neuroscience experiment at a research university, monitoring seizure activity in a clinical setting, or building a BCI application — the software layer is not where you want to cut corners or just default to whatever's familiar.
Spend time on this decision. Audit your current pipeline. Talk to collaborators. Try platforms before committing. And revisit the question as your research evolves — the right tool at year one of a project may not be the right tool at year five.
Looking for guidance on building or upgrading your EEG analysis pipeline? Connect with a neuroscience technology specialist who can evaluate your specific research needs and help you build a workflow that's rigorous, reproducible, and ready for where the field is heading.